Percolation Threshold on a Square Lattice
Manfred Schroeder touches on the topic of percolation a number of times in his encyclopaedic book on fractals (Schroeder, M. (1991) Fractals, Chaos, Power Laws: Minutes from an Infinite Paradise. Percolation has numerous practical applications, the most interesting of which (from my perspective) is the flow of hot water through ground coffee! The problem of percolation can be posed as follows: suppose that a liquid is poured onto a solid block of some substance. If the substance is porous then it is possible for the liquid to seep through the pores and make it all the way to the bottom of the block. Whether or not this happens is determined by the connectivity of the pores within the substance. If it is extremely porous then it is very likely that there will be an open path of pores connecting the top to the bottom and the liquid will flow freely. If, on the other hand, the porosity is low then such a path may not exist. Evidently there is a critical porosity threshold which divides these two regimes.
This situation can be modelled on a regular lattice where each of the lattice sites is either occupied (with probability p) or vacant (with probability 1-p). This is known as the site percolation problem. The related bond percolation problem can be posed in terms of whether or not the edges between neighbouring sites are open or closed. As we will see there is a well defined range of p (centred on a threshold value pc) for which the probability of percolation decreases rapidly from one to zero.
The percolation problem has been studied on a menagerie of lattice types but we will focus on the simplest, the square lattice.
Creating the Lattice
First we will load a few handy libraries and set up some constants.
library(ggplot2)
library(reshape2)
library(plyr)
EMPTY = 0
OCCUPIED = 1
FLOW = 2Next a function to generate and populate a lattice, which takes as arguments the width of the lattice and the occupation probability.
create.grid <- function(N, p) {
grid = matrix(rbinom(N**2, 1, p), nrow = N)
#
attributes(grid)$p = p
#
return(grid)
}
set.seed(1)
g1 = create.grid(12, 0.6)
g2 = create.grid(12, 0.4)Visualising the Lattice
g1 [,1] [,2] [,3] [,4] [,5] [,6] [,7] [,8] [,9] [,10] [,11] [,12]
[1,] 1 0 1 0 0 0 1 0 1 0 0 1
[2,] 1 1 1 1 0 1 1 1 1 1 1 0
[3,] 1 0 1 0 1 1 1 0 0 0 1 0
[4,] 0 1 1 1 0 1 0 1 0 0 1 1
[5,] 1 0 0 0 1 0 0 1 0 1 0 1
[6,] 0 0 1 0 1 1 1 1 1 1 1 1
[7,] 0 1 1 0 1 1 0 1 1 1 1 0
[8,] 0 0 1 1 1 0 0 1 0 1 1 1
[9,] 0 0 1 1 1 1 1 0 0 0 1 0
[10,] 1 1 1 0 1 0 0 0 1 1 1 0
[11,] 1 0 0 1 0 1 1 0 1 1 1 1
[12,] 1 1 0 1 1 0 1 0 1 0 1 1In the text representation of the lattice the occupied sites are represented by a 1, while a 0 indicates a vacant site.
visualise.grid <- function(g) {
N = nrow(g)
#
limits = c(0, N) + 0.5
#
ggplot(melt(g[nrow(g):1,], 1, varnames = c("row", "col"), value.name = "occupied")) +
geom_tile(aes(x = col, y = row, fill = factor(occupied))) +
geom_hline(yintercept = limits, size = 2) +
geom_vline(xintercept = limits, size = 2) +
xlim(limits) + ylim(limits) +
coord_fixed(ratio = 1) +
scale_fill_manual(values = c("0" = "white", "1" = "grey", "2" = "lightblue")) +
theme(panel.background = element_blank(),
axis.title = element_blank(),
axis.text = element_blank(),
axis.ticks = element_blank(),
legend.position = "none"
)
}
require(gridExtra)
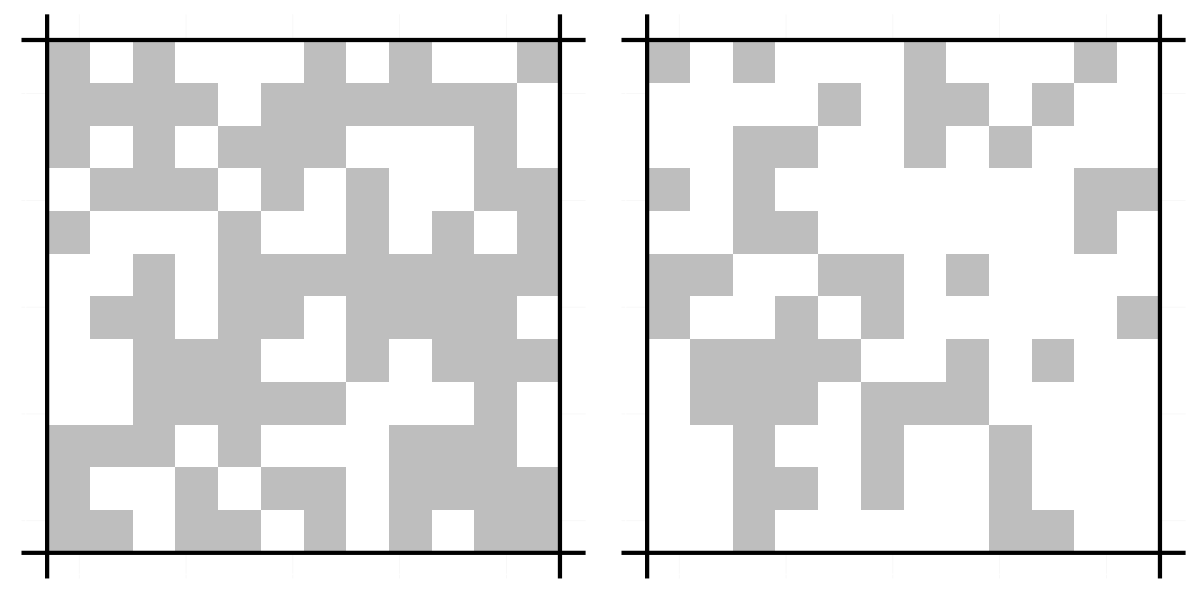
grid.arrange(visualise.grid(g1), visualise.grid(g2), ncol = 2)A graphical representation makes things a lot clearer.
Flow Through the Lattice
Now we need to check for flow through the lattice. We will use a recursive depth first search. Everything goes into a single function. The first argument is the grid. The second and third (optional) arguments are the row and column indices for a particular cell in the grid. If the latter arguments are omitted then the function loops through each of the cells in the top row of the grid. If a particular cell is specified, and it is not already occupied, then it is marked as being part of a flow path. The function then recursively considers each of the nearest neighbour cells.
flow <- function(g, i = NA, j = NA) {
# -> Cycle through cells in top row
#
if (is.na(i) || is.na(j)) {
for (j in 1:ncol(g)) {
g = flow(g, 1, j)
}
return(g)
}
#
# -> Check specific cell
#
if (i < 1 || i > nrow(g) || j < 1 || j > ncol(g)) return(g)
#
if (g[i,j] == OCCUPIED || g[i,j] == FLOW) return(g)
#
g[i,j] = FLOW
g = flow(g, i+1, j) # down
g = flow(g, i-1, j) # up
g = flow(g, i, j+1) # right
g = flow(g, i, j-1) # left
g
}The flow path can then be visualised.
grid.arrange(visualise.grid(flow(g1)), visualise.grid(flow(g2)), ncol = 2)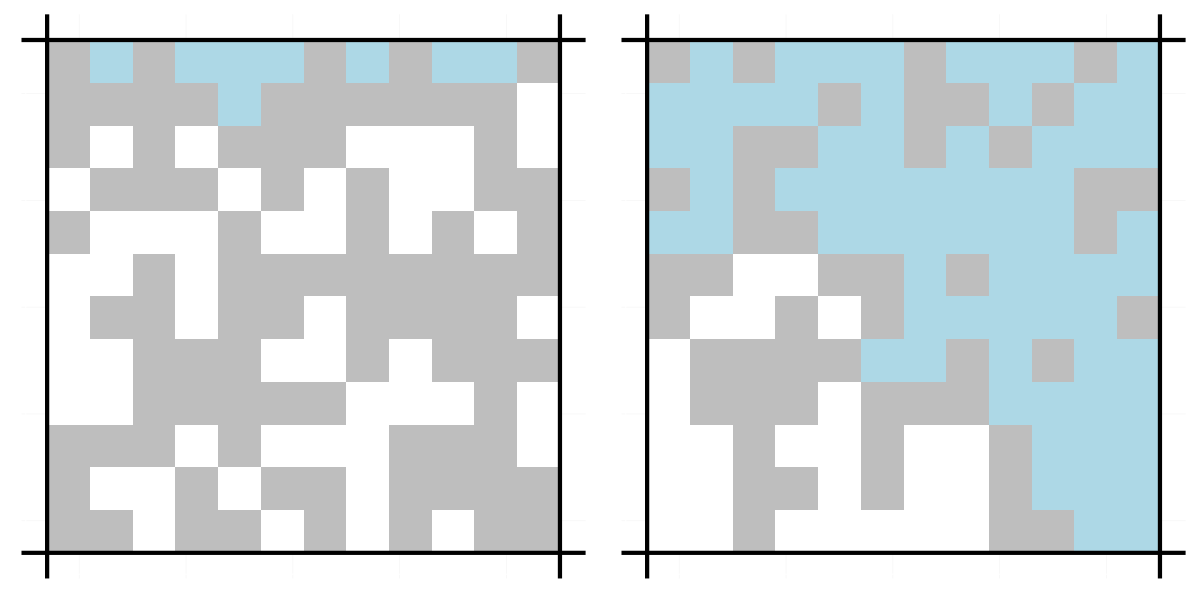
The grid on the left does not percolate while the one on the right does.
Checking for Percolation
Finally we can determine whether a given grid percolates by checking whether or not the flow makes it to the bottom of the grid.
# Check whether flow reaches to bottom of the lattice.
#
percolates <- function(g) {
g <- flow(g)
for (j in 1:ncol(g)) {
if (g[nrow(g), j] == FLOW) return(TRUE)
}
return(FALSE)
}
percolates(g1)[1] FALSEpercolates(g2)[1] TRUEFinding the Percolation Threshold
Finding the percolation threshold is a little numerically intensive, so it makes sense to take advantage of parallel execution.
library(foreach)
library(doMC)
registerDoMC(cores=4)We define some parameters for the calculation: the dimensions of the lattice and the number of replicates for each value of the occupation probability, p.
N = 25
REPLICATES = 1000Although we will be considering the full range of possible values for p, it makes sense to focus our attention on those values closer to the threshold. For each value of p we randomly generate a number of grids and check whether or not each one percolates.
pseq = unique(c(seq(0, 0.2, 0.1), seq(0.2, 0.6, 0.0125), seq(0.6, 1, 0.1)))
#
perc.occp = foreach(p = pseq, .combine=rbind) %dopar% {
data.frame(p, percolates = replicate(REPLICATES, percolates(create.grid(N, p))))
}Summary statistics are then compiled to give the percolation probability as a function of p.
perc.summary = ddply(perc.occp, .(p), summarize,
mean = mean(percolates),
sd = sd(percolates) / sqrt(length(percolates))
)A logistic model is fitted to these data and the threshold value for p is estimated.
logistic.fit = glm(mean ~ p, family=binomial(logit), data = perc.summary)
#
perc.summary$fit = logistic.fit$fitted
#
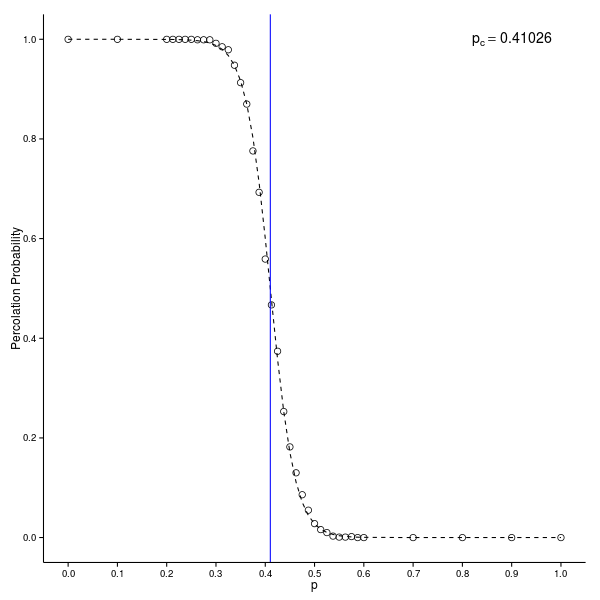
(pcrit = - logistic.fit$coefficients[1] / logistic.fit$coefficients[2])(Intercept)
0.410261 - pcrit(Intercept)
0.58974That compares pretty well with the reference results for the 44 square lattice.
Finally, all of this is put together in a plot.
ggplot(perc.summary, aes(x = p, y = mean)) +
geom_point(size = 3, shape = 21) +
geom_line(aes(x = p, y = fit), linetype = 2) +
geom_vline(xintercept = pcrit, color = "blue") +
annotate("text", label = sprintf("p[c] == %.5f", pcrit), x = 0.9, y = 1.0, parse = TRUE) +
scale_x_continuous(breaks = seq(0, 1, 0.1)) +
scale_y_continuous(breaks = seq(0, 1, 0.2)) +
ylab("Percolation Probability") +
theme_classic()